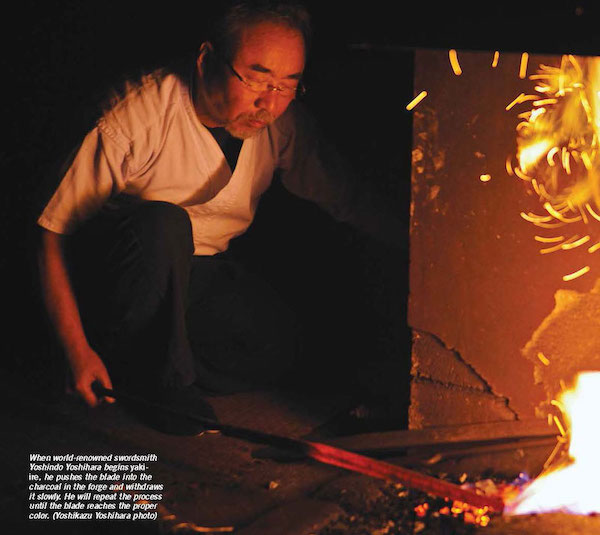
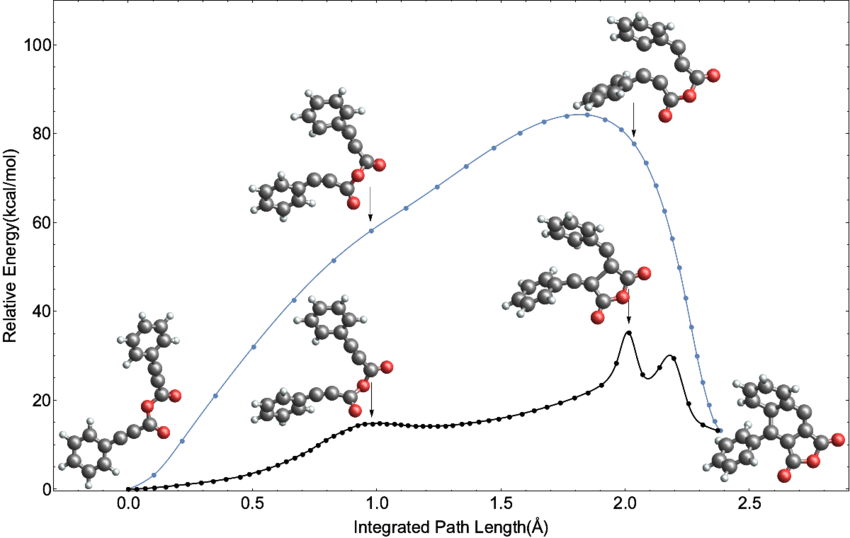
We that the decay follows very closely the expected exponential decay, but it is not exact, as the simulation has captured the random nature of the process.
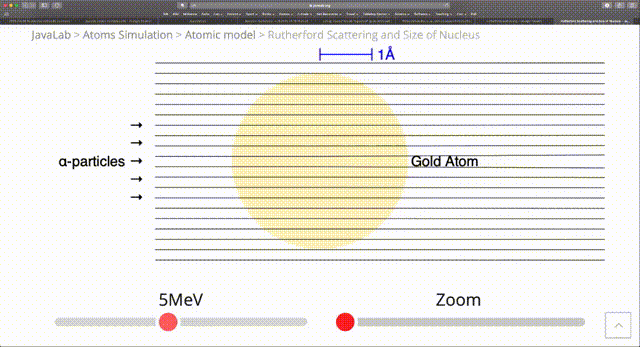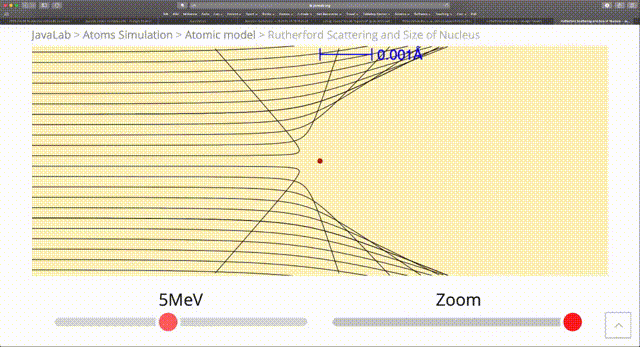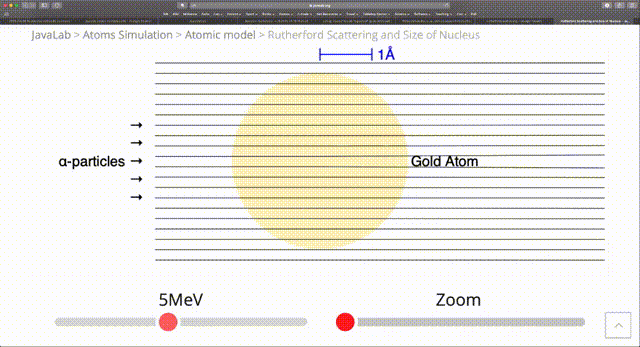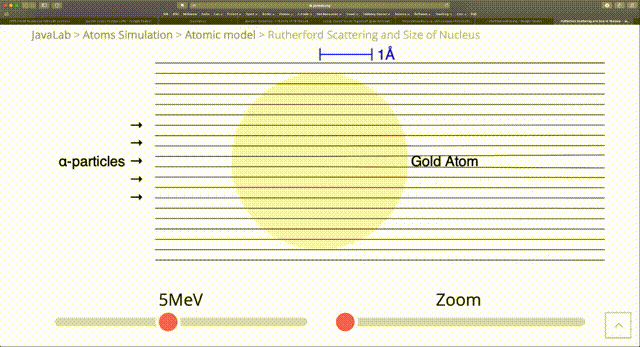
The standard RNG from numpy such as random provide a uniform distribution of random numbers. However, in computational physics there are many situations where we are modelling processes that happen with nonuniform probability. In the radioactive example we just considered, we had $N$ atoms with a half-life of $\tau$ and the probability $p(t)$ that a particular particle decays in time $t$ is $1 - 2^{-\frac{t}{\tau}}$. Then the probability that it decays in an infinitesimal time $dt$ is:
$$1 - 2^{-\frac{dt}{\tau}} = 1 - \exp\left(-\frac{dt}{\tau}\ln 2 \right) = \frac{\ln 2}{\tau}dt$$
ignoring terms in $(dt)^2$ or smaller. If we now focus on a single atom, we can ask what is the probability that it decays between times $t$ and $t+dt$. Firstly, it must survive until time $t$, which has a probability of $2^{-\frac{dt}{\tau}}$ and then decay in the following interval $dt$. The total probability $P(t)dt$ is then:
$$P(t)dt = 2^{-\frac{dt}{\tau}} \frac{\ln 2}{\tau}dt$$
This is an example of a nonuniform probability distribution, as the decay times are proportional to $2^{-\frac{t}{\tau}}$, so that earlier decay times are more likely than later ones, although all are possible. The point of this, is that we now have a much more efficient method of solving the radioactive decay problem. Instead of running through every single atom at every timestep, we could just draw $N$ random numbers from this nonuniform probability distribution and count how many decay before a give time. To do this, we need a function that generates nonuniform random numbers and this is usually done by applying a transformation method to a standard RNG.
Suppose we have a source of random floating point numbers $z$ from a distribution $q(z)$ with a probability of getting a number between $z$ and $z+dz$ being $q(z)dz$. We also have a function $x=x(z)$, which gives us another random number from $z$, but with another distribution $p(x)$, different from $q(z)$. We want to define $x(z)$ such that $x$ has the distribution we want. The probability of generating a value of $x$ between $x$ and $x + dx$ is by definition equal to the probability of generating a value of $z$ in the equivalent interval:
$$p(x)dx = q(z)dz$$
Let's assume we are using the numpy function random, so what we have an equal probability of getting a number from zero to one. This means $q(z)=1$ from zero to one and is zero everywhere else. Then we can integrate the probabilities as follows:
$$\int_{-\infty}^{x(z)} p(x^\prime)dx^\prime = \int_0^z dz^\prime = z$$
If we can integrate the function on the left, we have an equation that must be solved by the value of $x(z)$, and if we can solve that, we get the function that we want. As an example, let us consider the set of random real numbers $x$ in the interval from zero to infinity for the exponential probability distribution:
$$p(x) = \mu e^{-\mu x}$$
using the same radioactive decay distribution $\mu = \ln 2/\tau$, which looks suspiciously normalized and integrable...
$$\mu \int_{0}^{x(z)} \text{e}^{-\mu x^\prime}dx^\prime = 1 - \text{e}^{-\mu x} = z$$
then:
$$x = -\frac{1}{\mu} \ln(1-z)$$
So we just need to generate uniform random numbers $z$ from zero to one and then feed them into this equation to get exponentially distributed $x$ values. Note that in principle whether you use $\ln(1-z)$ or $\ln(z)$ should not matter in terms of randomness, but in practice, your uniform distribution will generally either include 0 (numpy.random) or 1, not both (generators are usually in the half-open interval), and you should choose to avoid the discontinuity in the simulations. You can usually specify this when calling the RNG.